Introduction
Angelman Syndrome is a rare neurological disorder that affects roughly 1 in every 15,000 people. Classic symptoms of Angelman Syndrome include developmental delay, speech and balance impairments, excessive excitability, hand flapping, and trouble with sleep. However, only some cases of Angelman Syndrome display additional symptoms, such as epilepsy. It is known that Angelman Syndrome patients with epilepsy have a specific mutation in the gene, SNRPN, but it is unknown what role this mutation plays in disease.
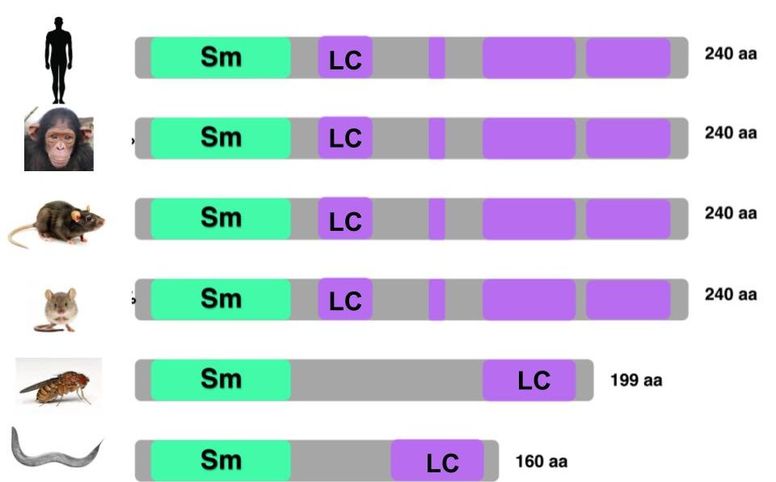
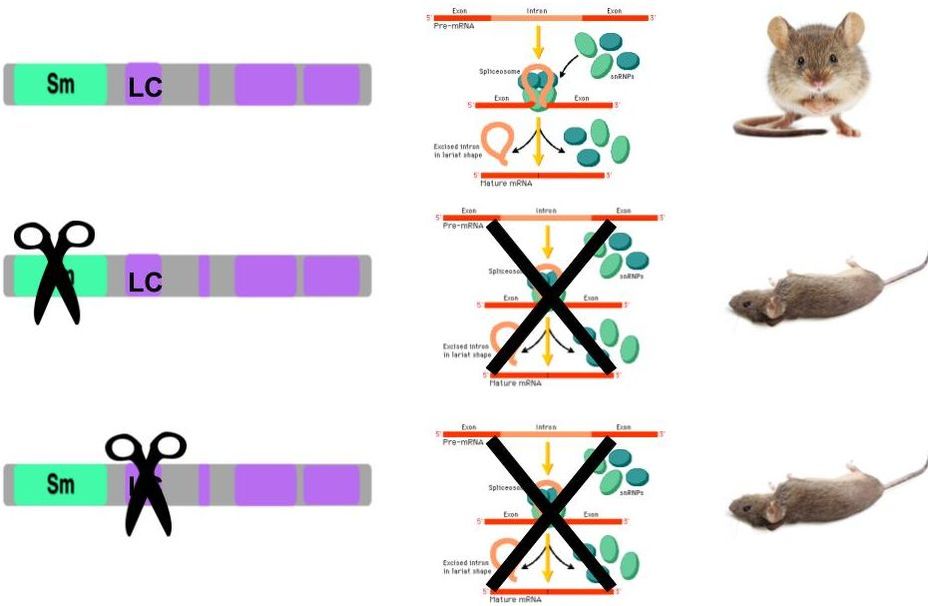
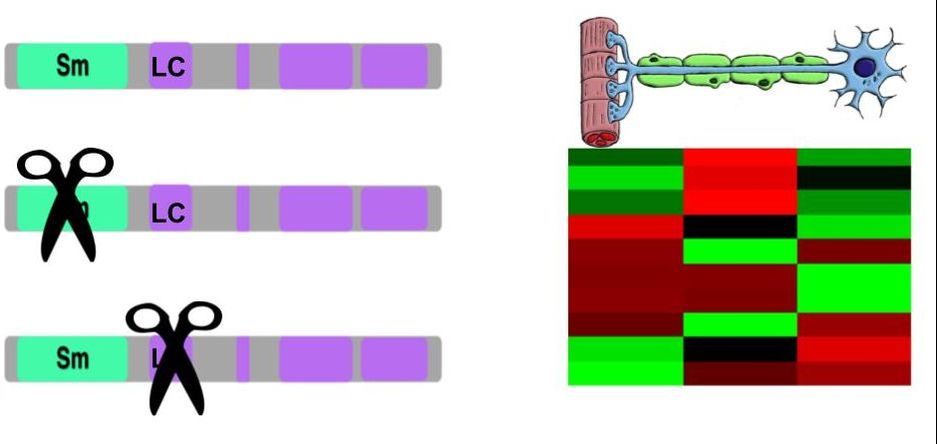
SNRPN is a protein conserved across a diverse array of species, and consists of an Sm domain, which is implicated in RNA binding, and multiple Low Complexity (LC) domains of unknown function (Figure 1).
SNRPN is a protein conserved across a diverse array of species, and consists of an Sm domain, which is implicated in RNA binding, and multiple Low Complexity (LC) domains of unknown function (Figure 1).
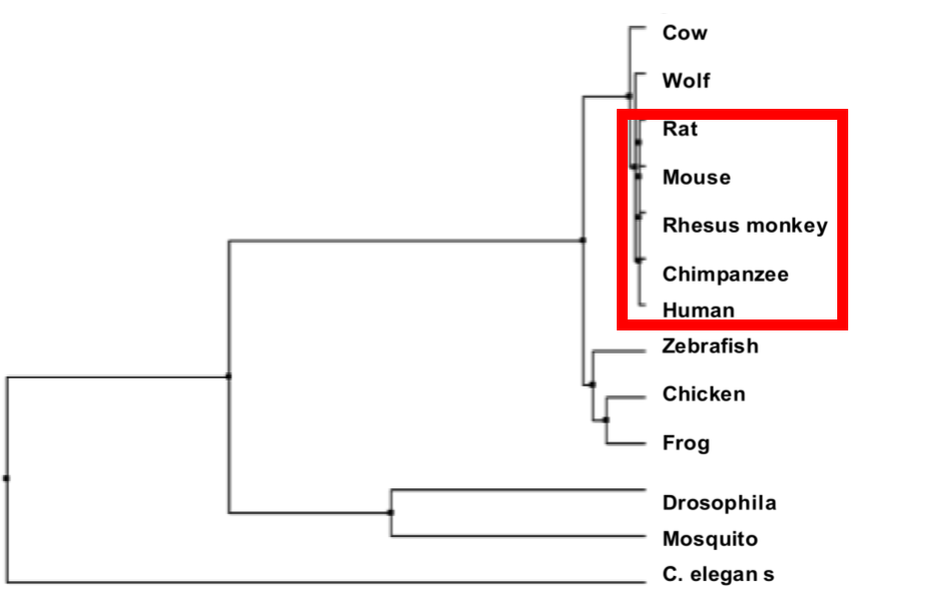
Figure 1. Left: SNRPN homologs in different species. Right: Phyolgenetic relationships of SNRPN homologs. Red box indicates species with 100% similarity between homolog sequences.
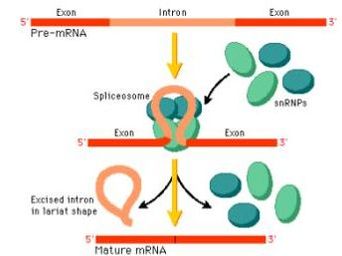
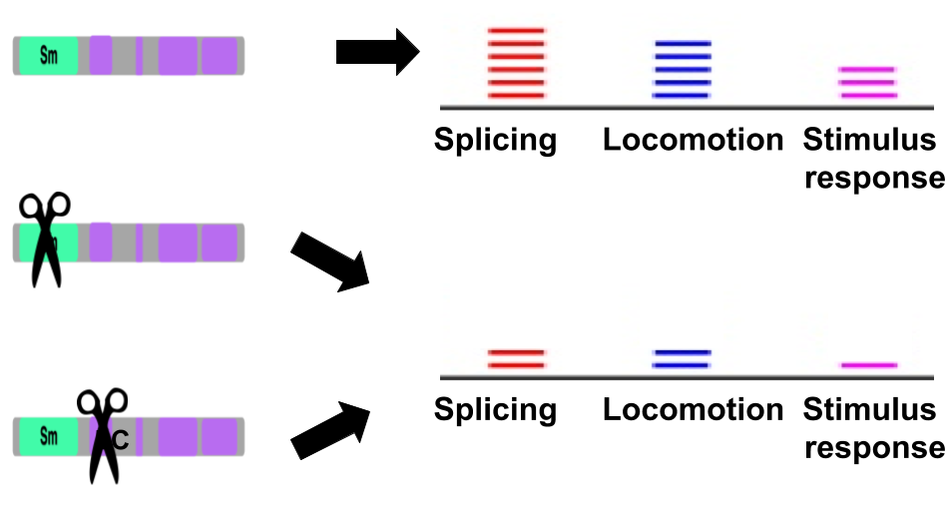
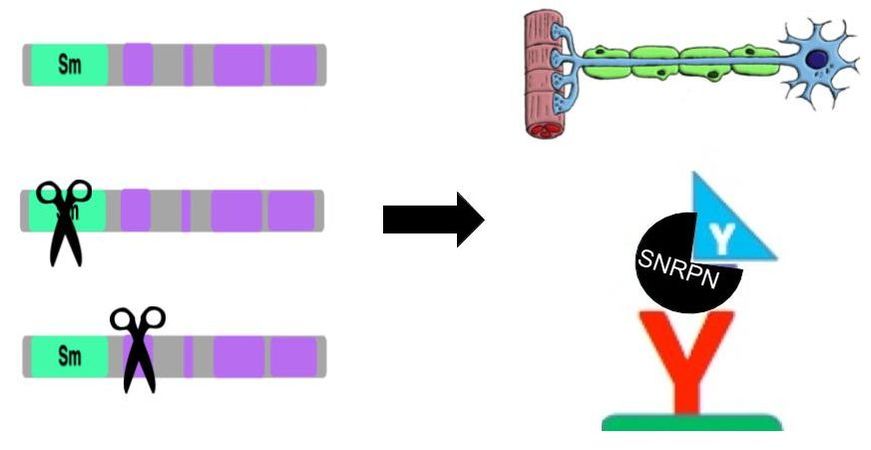
Through the Gene Ontology database, I found that SNRPN primarily functions as an RNA processing enzyme that is present at high levels in neurons. Its molecular function is RNA binding through the Sm domain, its biological process is RNA processing via the spliceosome, and its cellular component is the nucleoplasm.
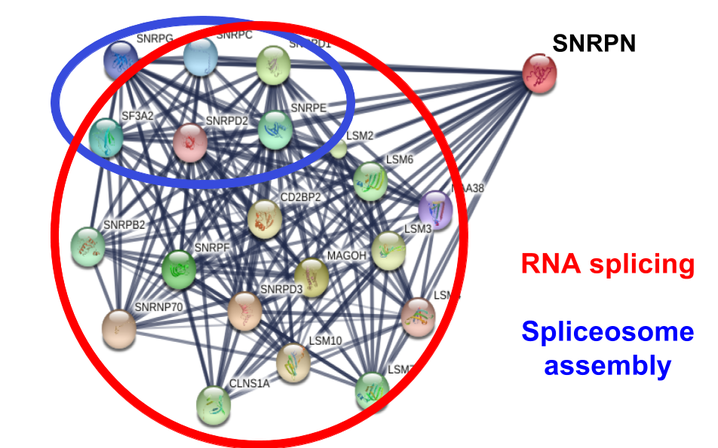
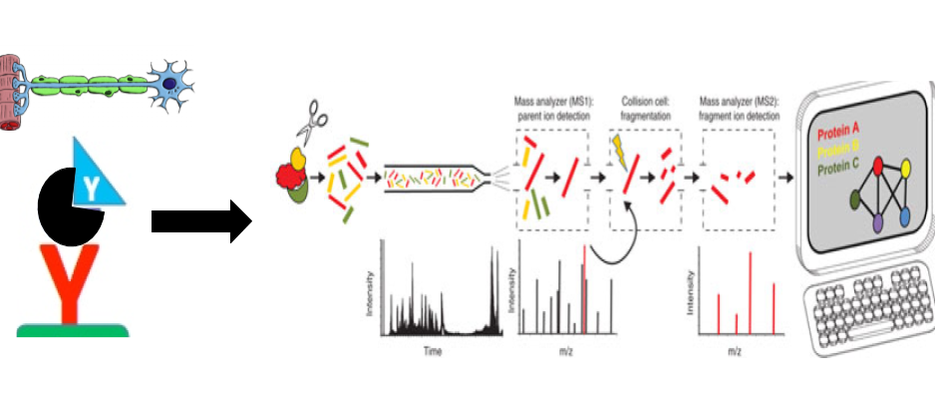
As a component of the spliceosome, SNRPN interacts with an extensive number of proteins (Figure 5). This includes not only other direct components of the spliceosome, but also scaffolding proteins and regulatory proteins. Because of the many components involved in RNA processing and spliceosome assembly, it can be difficult to determine what specific proteins are critical for proper function. With this in mind, SNRPN function in neurons can be analyzed in the context of epilepsy in cases of Angelman Syndrome.
Figure 5. Human SNRPN and its protein interaction network.
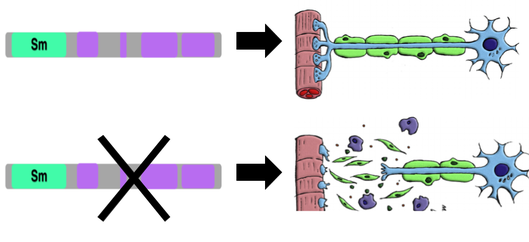
It is known that improper splicing in neurons can lead to neurodegeneration, which may result in downstream epilepsy. Although epileptic Angelman Syndrome patients have specific SNRPN mutations, it is unknown what role SNRPN plays in proper neuronal function.
My goal is to determine what is the role of RNA processing in neuronal function.
My hypothesis is that SNRPN is critical for RNA processing, and SNRPN dysfunction leads to epilepsy.
My long-term goal is to determine if defects in SNRPN RNA processing is responsible for other phenotypic effects in patients with Angelman Syndrome.
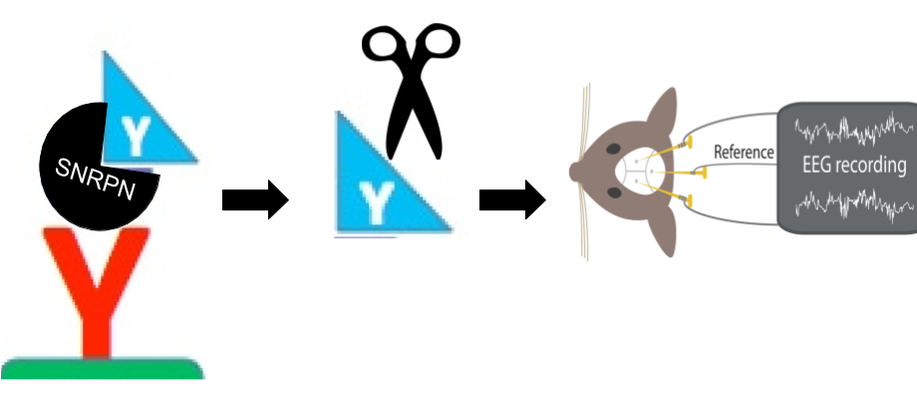
To investigate this goal and hypothesis, I have proposed a series of experiments, or Specific Aims, that will address different components of the role of SNRPN in RNA processing, and detect connections to epilepsy. All of these experiments will be conducted in mice because not only is the mouse SNRPN homolog 100% identical to human SNRPN, but also the SNRPN mutant phenotype in mice is epilepsy. Furthermore, epilepsy can be easily quantified and detected in mice using an EEG test (Figure 6).
My goal is to determine what is the role of RNA processing in neuronal function.
My hypothesis is that SNRPN is critical for RNA processing, and SNRPN dysfunction leads to epilepsy.
My long-term goal is to determine if defects in SNRPN RNA processing is responsible for other phenotypic effects in patients with Angelman Syndrome.
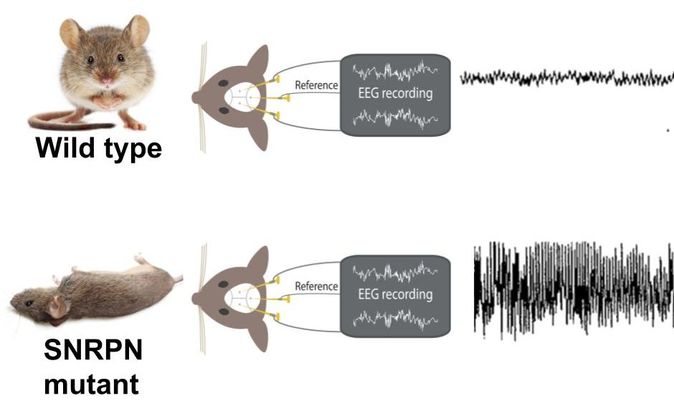
To investigate this goal and hypothesis, I have proposed a series of experiments, or Specific Aims, that will address different components of the role of SNRPN in RNA processing, and detect connections to epilepsy. All of these experiments will be conducted in mice because not only is the mouse SNRPN homolog 100% identical to human SNRPN, but also the SNRPN mutant phenotype in mice is epilepsy. Furthermore, epilepsy can be easily quantified and detected in mice using an EEG test (Figure 6).
Figure 6. EEG epilepsy test comparisons between wildtype/non-epileptic mice (top) and SNRPN mutant/epileptic mice (bottom>
Specific Aims
Future Directions
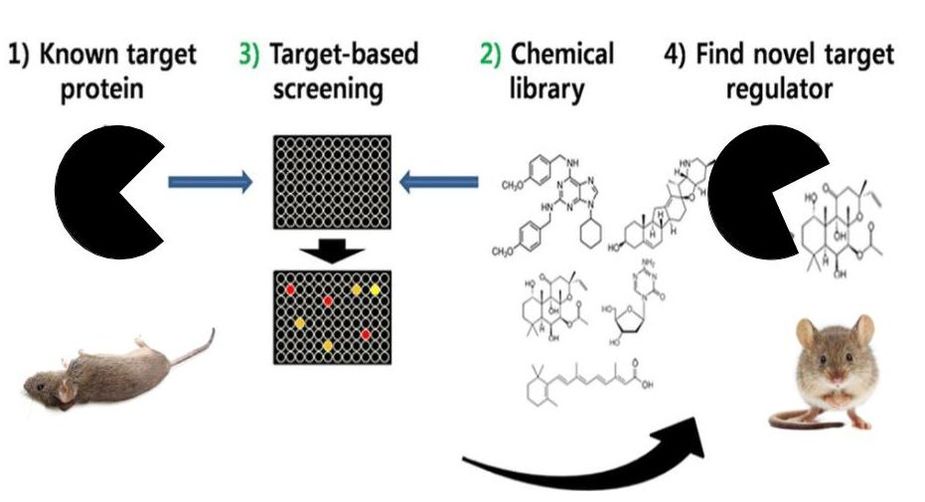
Going forward, I would design an experiment to determine if SNRPN mutants can be rescues. To asses this, I would perform a chemical screen on different SNRPN domain mutants to identify any potential small molecules that ameliorate epileptic phenotypes in mutant mice (Figure 13).
Figure 13. Chemical screen identifying small molecules with therapeutic potential.
Talk Downloads
| fischer_finalpresentation_27april2017.pptx.pdf | |
| File Size: | 2257 kb |
| File Type: | |
| fischer_presentationdraft2_4-4-17.pptx | |
| File Size: | 6314 kb |
| File Type: | pptx |
| fischerjulie_21feb2017_draftpresentation__1_.pdf | |
| File Size: | 341 kb |
| File Type: | |
References
[1] Angelman Syndrome: Genetics Home Reference
<https://ghr.nlm.nih.gov/condition/angelman-syndrome>
[2] Li, H., Zhao, P., Xu, Q., Shan, S., Hu, C., Qiu, Z., & Xu, X. (2016). The autism-related gene SNRPN regulates cortical and spine development via controlling nuclear receptor Nr4a1. Scientific Reports, 6, 29878. doi:10.1038/srep29878
<https://www.ncbi.nlm.nih.gov/pubmed/1533223>
[3] Neuropathology: An illustrated interactive course for medical students and residents <http://neuropathology-web.org/chapter9/chapter9hAtaxia.html>
[4] Gallo, J. . (2005). The role of RNA and RNA processing in Neurodegeneration. Journal of Neuroscience, 25(45), 10372–10375. doi:10.1523/jneurosci.3453-05.2005
< http://www.jneurosci.org/content/25/45/10372 >
[5] Battaglia, A., Gurrieri, F., Bertini, E., Bellacosa, A., Pomponi, M. G., Paravatou-Petsotas, M., . . . Neri, G. (1997). The inv dup(15) syndrome: A clinically recognizable syndrome with altered behavior, mental retardation, and epilepsy. Neurology, 48(4), 1081-1086. doi:10.1212/wnl.48.4.1081
<https://www.ncbi.nlm.nih.gov/pubmed/9109904>
[6] Johnstone, K. A. (2005). A human imprinting centre demonstrates conserved acquisition but diverged maintenance of imprinting in a mouse model for Angelman syndrome imprinting defects. Human Molecular Genetics, 15(3), 393-404. doi:10.1093/hmg/ddi456
<https://www.ncbi.nlm.nih.gov/pubmed/16368707?dopt=Abstract>
<https://ghr.nlm.nih.gov/condition/angelman-syndrome>
[2] Li, H., Zhao, P., Xu, Q., Shan, S., Hu, C., Qiu, Z., & Xu, X. (2016). The autism-related gene SNRPN regulates cortical and spine development via controlling nuclear receptor Nr4a1. Scientific Reports, 6, 29878. doi:10.1038/srep29878
<https://www.ncbi.nlm.nih.gov/pubmed/1533223>
[3] Neuropathology: An illustrated interactive course for medical students and residents <http://neuropathology-web.org/chapter9/chapter9hAtaxia.html>
[4] Gallo, J. . (2005). The role of RNA and RNA processing in Neurodegeneration. Journal of Neuroscience, 25(45), 10372–10375. doi:10.1523/jneurosci.3453-05.2005
< http://www.jneurosci.org/content/25/45/10372 >
[5] Battaglia, A., Gurrieri, F., Bertini, E., Bellacosa, A., Pomponi, M. G., Paravatou-Petsotas, M., . . . Neri, G. (1997). The inv dup(15) syndrome: A clinically recognizable syndrome with altered behavior, mental retardation, and epilepsy. Neurology, 48(4), 1081-1086. doi:10.1212/wnl.48.4.1081
<https://www.ncbi.nlm.nih.gov/pubmed/9109904>
[6] Johnstone, K. A. (2005). A human imprinting centre demonstrates conserved acquisition but diverged maintenance of imprinting in a mouse model for Angelman syndrome imprinting defects. Human Molecular Genetics, 15(3), 393-404. doi:10.1093/hmg/ddi456
<https://www.ncbi.nlm.nih.gov/pubmed/16368707?dopt=Abstract>