This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are Model Organisms?
Model organisms are non-human creatures used to study human processes. Model organisms are selected based on their similarity to the human system that is being studied, as well as for their lower-maintenance qualities and ease of studying. Some of the benefits of conducting a study with a model organism include lower prices, quicker reproduction rates, and ability to conduct more trials than in human studies.
What Model Organisms can be Used to Study SNRPN?
Mice are an appropriate model for studying the function of SNRPN in humans. SNRPN in mice is 100% similar to the human SNRPN homolog, therefore the structure and interactions should accurately reflect that of human SNRPN [1]. In mice, the SNRPN mutant phenotype is epilepsy [2]. Furthermore, epilepsy in mice can be easily quantified using an EEG test.
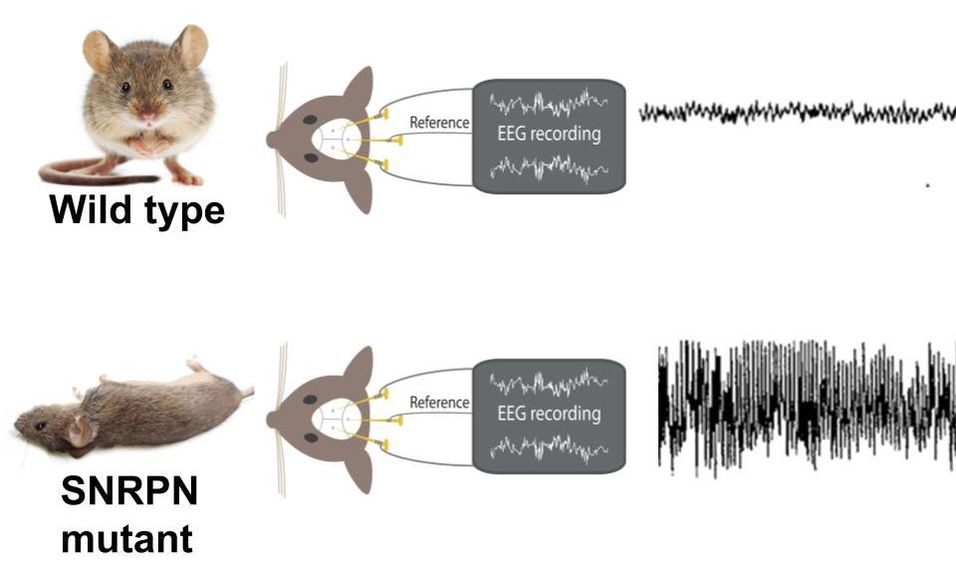
Figure 1. EEG recordings comparing wildtype/non-epileptic mice to SNRPN mutant/epileptic mice.
What are the benefits of using mice as a model organism?
Aside from the mouse SNRPN mutant phenotype discussed above, mice also offer additional benefits as a model organism. Mice display a similar physiological makeup compared to humans, and are relatively feasible to care for [3]. Furthermore, they are inexpensive and have a high reproductive rate such that many experimental replicates can be conducted. Mice are also an appropriate model to study SNRPN in the context of epilepsy because the human and mouse brain are 90% identical in their composition [4].
What are the mutant phenotypes in model organisms?
The mutant version of SNRPN homologs in different model organisms can indicate what types of biological processes SNRPN is important for. In addition, the mutant phenotypes can provide helpful information when selecting what model organisms to use.
Mouse SNRPN homolog: Snrpn
Mutant phenotype: epilepsy
Mutant phenotype: epilepsy
C. elegans SNRPN homolog: snr-2
Mutant phenotype: abnormal nuclear and nucleolus morphology [5]. Human SNRPN localizes in the nucleolus and processes RNA here so that is can be transported to the cytoplasm for translation.
Mutant phenotype: abnormal nuclear and nucleolus morphology [5]. Human SNRPN localizes in the nucleolus and processes RNA here so that is can be transported to the cytoplasm for translation.
Fruit Fly SNRPN homolog: SMB
Mutant phenotype: tau toxicity [6]. In humans, tau toxicity is implicated in neurodegenerative disease, which also can lead to downstream epilepsy.
Mutant phenotype: tau toxicity [6]. In humans, tau toxicity is implicated in neurodegenerative disease, which also can lead to downstream epilepsy.
References
[1] NCBI: SNRPN Mus musculus <https://www.ncbi.nlm.nih.gov/protein/133725810>
[2] Johnstone, K. A. (2005). A human imprinting centre demonstrates conserved acquisition but diverged maintenance of imprinting in a mouse model for Angelman syndrome imprinting defects. Human Molecular Genetics, 15(3), 393-404. doi:10.1093/hmg/ddi456 <https://www.ncbi.nlm.nih.gov/pubmed/16368707?dopt=Abstract>
[3] Yg Topics: Why use the mouse in research? <http://www.yourgenome.org/facts/why-use-the-mouse-in-research>
[4] Allen Brain Atlas Data Portal <http://www.brain-map.org/>
[5] Worm Base <http://www.wormbase.org/species/all/phenotype/WBPhenotype:0001028#0--10>
[6] Fly Base <Fhttp://flybase.org/reports/FBgn0262601.html>
[2] Johnstone, K. A. (2005). A human imprinting centre demonstrates conserved acquisition but diverged maintenance of imprinting in a mouse model for Angelman syndrome imprinting defects. Human Molecular Genetics, 15(3), 393-404. doi:10.1093/hmg/ddi456 <https://www.ncbi.nlm.nih.gov/pubmed/16368707?dopt=Abstract>
[3] Yg Topics: Why use the mouse in research? <http://www.yourgenome.org/facts/why-use-the-mouse-in-research>
[4] Allen Brain Atlas Data Portal <http://www.brain-map.org/>
[5] Worm Base <http://www.wormbase.org/species/all/phenotype/WBPhenotype:0001028#0--10>
[6] Fly Base <Fhttp://flybase.org/reports/FBgn0262601.html>